INTRODUCTION
Since December 2019, an unknow pneumonia associated with a novel human coronaviruses, the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), named by the International Committee on Taxonomy of Viruses (ICTV) and named as coronavirus disease 2019 (COVID-19) by World Health Organization (WHO), emerged in Wuhan, China 1-3. The ongoing outbreak of COVID-19 and its rapid transmission to 25 other countries across the world resulted in a global health emergency declared by WHO on January 30, 2020 4. On March 11, 2020, the epidemic spread rapidly worldwide and was characterized as a pandemic by WHO 5. The COVID-19 pandemic is sorely testing health care systems around the world and is considered as the major health emergency in a century 6.
Coronaviruses (CoVs) are a family of enveloped viruses containing single-stranded positive sense RNA (+ssRNA) that belongs to Coronaviridae family of the Orthocoronavirinae subfamily of the Nidovirales order 7,8. CoVs are genotypically and serologically divided into four genera: Alphacoronavirus, Betacoronavirus, Gammacoronavirus, and Deltacoronavirus 3. Based on phylogenetic and genomic relationships, SARS-CoV-2 belongs to genera Betacoronavirus 9. CoVs virions are lipid enveloped and spherical with a size of approximately 100-120 nm; composed of a nucleocapsid and an external envelope. The nucleocapsid houses and protects the RNA genome associated with the nucleocapsid (N) protein 10. The external envelope is composed of major structural proteins (Spike (S), Membrane (M) and Envelope (E) proteins) and accessory proteins 11. SARS-CoV-2 is known to have at least seven accessory proteins (Hemagglutinin-esterase (HE), 3a, 6, 7a, 7b, 8, and 10) 12 (Figure 1). The S protein is important for attachment to the host cells, the M and E proteins are necessary for virus assembly, the N protein is involved in the transcription and replication of the viral RNA and packaging of the genome into new virions 11.
SARS-CoV-2 is closely related to other human CoVs that caused zoonotic epidemics, these are Severe Acute Respiratory Syndrome (SARS-CoV) and Middle Eastern Respiratory Syndrome (MERS-CoV), which caused local outbreaks in 2003 and 2012, respectively 13. Nonetheless, among all known CoVs, SARS-CoV-2 is more similar to bat coronavirus RaTG13. The genome sequence of SARS-CoV-2 is 98% identical with RaTG13, suggestting that bat may be a natural host of SARS-CoV-2 14. CoVs are known to infect and cause disease animals and humans. Among human CoVs, four of them (229E, NL63, OC43 and HKU1) infect typically only the upper respiratory tract and cause relatively minor symptoms. SARS-CoV, MERS-CoV and SARS-CoV-2 predominantly infect the lower respiratory tract and can cause pneumonia, which can be fatal 15,16.
Clinical Features of COVID-19 and Cytokine Storm Syndrome
The rapid spread of COVID-19 is explained in part due to nature of viral transmission. The median incubation time of SARS-CoV-2 infection ranges from four to seven days creating a large window of time for viral transmission. Many patients are completely asymptomatic or have few symptoms yet are fully capable of transmitting the virus 17. The main clinical symptoms typically include fever, cough, anosmia, dysgeusia, fatigue, myalgia, and gastrointestinal symptoms such as, loss of appetite, nausea and diarrhoea 6. Most patients with COVID-19 exhibit mild to moderate symptoms, but approximately 15% progress to severe pneumonia and 5% eventually develop acute respiratory distress syndrome (ARDS), septic shock and/or multiple organ failure, conditions that are the major threat and are the major cause of death 18. In severe patients, respiratory support such as, general oxygen therapy, high-flow oxygen, noninvasive ventilation, and invasive mechanical ventilation is essential to relieve the symptoms and save their lives 19. In addition, it has been indicated that 25.9% of patients with SARS-CoV-2 pneumonia require intensive care unit (ICU) admission and 20.1% develop ARDS. The mortality rate appears to be around 2% 20.
COVID-19 morbidity and mortality have been linked to elderly age and comorbidities, such as hypertension, chronic obstructive pulmonary disease (COPD), diabetes, and cardiovascular disease, leading to a poorer outcome to the viral infection and, more frequently hospitalization and ICU admittance. Among such individuals, cancer patients represent a subgroup at high risk of developing severe complications from SARS-CoV-2 infection 21. History of smoking also should be considered as a predisposing factor in these patients with increased susceptibility to infection with COVID-19 22. These individuals are considered at high-risk of experiencing one of the most severe complication of COVID-19, that is, the cytokine storm syndrome 23. The effective antiviral responses of innate and adaptive immunity include the production of several pro-inflammatory cytokines and the activation of helper T cells (CD4+) and cytotoxic T cells (CD8+). However, the tissue injury caused by the virus could induce the exaggerated production of pro-inflammatory cytokines and the recruitment of pro-inflammatory macrophages and granulocytes, resulting in the cytokine storm syndrome 14,24. Clinical findings have shown exuberant inflammatory responses during SARS-CoV-2 infection, further resulting in uncontrolled pulmonary inflammation. A rapid viral replication may cause massive epithelial and endothelial cell death and vascular leakage, triggering the production of exuberant pro-inflammatory cytokines and chemokines, such as IL2, IL-6, IL7, IL10, GCSF, IP10, MCP1, MIP1A, and TNFα 25,26. In severe infections, analyzes have shown that the plasma levels of IL-2, IL-6, IL-8, IL-10, and TNF-α are higher than in non-severe infections. Furthermore, the plasma levels of IL-2, IL-7, IL-17, IL-10, MCP-1, MIP-1A and TNF-α in ICU patients are higher than in non-ICU patients 1,14. In addition, In patients with severe COVID-19, but not in patients with mild disease, lymphopenia is a common feature, with drastically reduced numbers of helper T (CD4+) cells, cytotoxic T (CD8+) cells, B cells and natural killer (NK) cells 19. Patients with severe SARS-CoV-2 infection have shown reduction and hyperactivation of peripheral helper T (CD4+) cells and cytotoxic T (CD8+) cells 23. Helper T cells (CD4+) are remarkably lower in severe patients than non-severe patients. These conditions could be a critical factor for severity and mortality of the disease 26.
Absence of a Specific Therapeutic Treatment Against SARS-CoV-2 Infection
Currently, there are no licensed approved treatments available for any coronavirus disease, including SARS-CoV-2 27. Treatment strategies for the disease include drugs uncommonly used in clinical practice 28. In efforts to reduce the mortality related to severe COVID-19, attempts have been made to design therapies that limit virus replication or modulate the host immune response 29. Antivirals are a class of small molecules that function as inhibitors of one or more stages of a virus life cycle such as, viral entry or replication. Because of similarities between different virus replication mechanisms, some antivirals can be repurposed against various viral infections 13. Current treatment options are mainly based on previous experience showing clinical benefits in treating influenza, Ebola, MERS-CoV, SARS-CoV, or other RNA and DNA viruses 30.
On the other hand, the most common therapeutic options for viral infections are promoting durable cellular and humoral immunity for the uninfected population via vaccination 17. However, a vaccine development can will take years before to test and reach in a large population. An optimistic estimate for any vaccine trial is at least 12-18 months 31,32.
For the prevention and treatment of COVID-19 disease, numerous clinical trials are ongoing to evaluate novel vaccine candidates and drug repurposing strategies against SARS-CoV-2 infection. Nevertheless, it is unclear how long these studies will take to establish efficacy and safety 32.
The objective of this article is to overview the different therapeutic agents that are being proposed and tested for the treatment of COVID-19 disease based on the blocking and inhibition of the molecular mechanisms of the viral life cycle of SARS-CoV- 2 infection, and for the treatment of the cytokine storm syndrome.
METHODS
A narrative review was performed by searching in the PubMed database with the search algorithm: (((Therapeutics AND Viral Life Cycle OR Cytokine Storm Syndrome OR ACE2 OR Spike Protein OR 3-chymotrypsin-like protease OR papain-like protease OR RNA-dependent RNA polymerase) OR Immunotherapy) AND (COVID-19 OR SARS-CoV-2)). All types of articles were included with the filters: Free full text, Full text, published in the last 1 year and in English. Articles were selected according to the objective of this review.
RESULTS
Viral Life Cycle of SARS-CoV-2
Similar to SARS-CoV, SARS-CoV-2 utilizes host receptor, angiotensin-converting enzyme 2 (ACE2), for its attachment and entry 33 ACE2 is a zinc metalloprotease that contributes to regulation of blood pressure by converting angiotensin II to angiotensin 1-7) (26,33,34. ACE2 is expressed in respiratory tract, heart, kidney, stomach, bladder, esophagus, and intestine, and mainly expressed in a small subset of cells called alveolar type II cells in the lung 2,26,35. Also, ACE2 is present in oral cavity in epithelial cells of salivary glands and epithelial cells of the tongue 36-38.
The S protein of CoVs facilitates viral entry into target cells 39. The S protein comprises two subunits, S1 and S2 40. The S1 subunit consists of an amino-terminal domain and a receptor- binding domain (RBD). S2 subunit allows fusion of viral and cellular membranes 39. The RBD in S1 subunit binds to ACE2 as its host cell target receptor and initiates the infection process 41. Computational modelling and biophysical measurements indicate that the SARS-CoV-2 RBD binds to ACE2 with higher affinity than that of SARS- CoV 15. Cryogenic Electron Microscopy (Cryo-EM) studies showed that SARS-CoV-2 had a ten-fold higher affinity for ACE2 compared to SARS-CoV. 42. This high affinity of the S protein of SARS-CoV-2 for human ACE2 may lead to the great human-to-human transmission 41. RBD binding to ACE2 triggers endocytosis of the SARS-CoV-2 virion and exposes it to endosomal proteases 15. Entry requires activation of S protein by transmembrane serine protease 2 (TMPRSS2) via proteolytic cleavage 39,43. This cleavage occurs in 2 different positions of S2, allowing the separation between RBD of S1 with ACE2, and the subsequent fusion of both membranes, facilitating the viral entry by endocytosis 9,44. After membrane fusion between the virus and the host cell, the viral genome reaches the cytoplasm 45.
The SARS-CoV-2 genome is a single-strand positive-sense RNA (+ssRNA) of approximately 30 kilobases (kb) 46. This genome acts as a typical mammalian messenger RNA (mRNA) due to its positive-sense and 5′-terminal cap structure and a poly(A) sequence at the 3′-end 47. Nonetheless, unlike most mammalian mRNA, this carries at least six open reading frames (ORFs) between the 5′-terminal and 3′-end 9. All CoVs have two large ORFs, (ORF1a and ORF1b), that occupy the 5′-terminal two-thirds of the genome and encode the viral replicase gene 9,47. At the 3’-end, one-thirds of the genome encodes for main structural proteins (Spike, M, E and N) and accessory proteins (HE, 3a, 6, 7a, 7b, 8, and 10) 9,12.
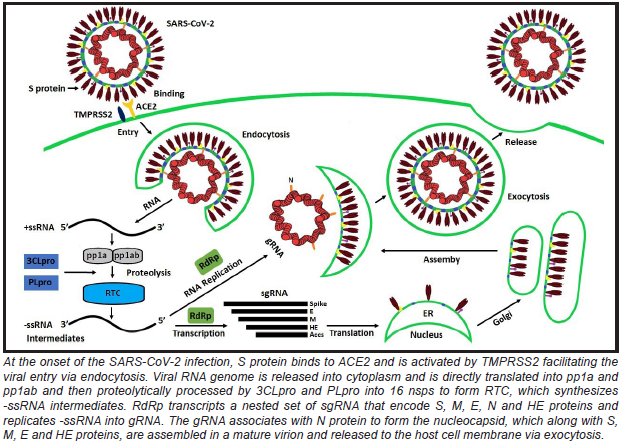
In the cytoplasm, the viral replicase gene is directly translated into two large polyproteins (pp1a and pp1ab) 9. pp1a and pp1ab are proteolytically processed by protease enzymes namely 3-chymotrypsin-like protease (3CLpro), and viral papain-like protease (PLpro) 48,49 into 16 non-structural proteins (nsps) 50-53. Some of these nsps are involved in ensuring viral RNA is translated efficiently without host interference and in the production of mature virions 54. The rest of the nsps are necessary to form the replicase-transcriptase complex (RTC) in double-membrane vesicles 2,9,49. The RTC is required to replicate viral RNA through single-strand negative-sense RNA (-ssRNA) intermediates 54. The nsp12, which harbor RNA-dependent RNA polymerase (RdRp) activity, takes the viral +ssRNA genome as a template for replication and transcription. The -ssRNA intermediates are generated to serve as the templates for the synthesis of positive-sense genomic RNA (gRNA) and a nested set of subgenomic RNAs (sgRNAs) 12. The sgRNAs encode for main structural proteins (Spike, M, E and N) and accessory proteins (HE, 3a, 6, 7a, 7b, 8, and 10) 2,9,12. The gRNA recently synthesized, associates with N protein and form the nucleocapsid in the cytoplasm. Main structural proteins (Spike, M and E) and accessory proteins are synthesized in endoplasmic reticulum (ER) and then, transported to Golgi. These proteins, along with the nucleocapsid, are assembled in a mature virion and released to the host cell membrane via exocytosis 2. A schematic diagram of the viral life cycle of SARS-CoV-2 is shown in (Figure 2).
Blocking the Spike (S) protein by Neutralizing Antibodies (NAbs)
In the absence of a vaccine, passive immunotherapy becomes an option for treating the SARS-CoV-2 infection. Passive immunotherapy involves the administration NAbs against an etiologic agent previously generated by an infected individual to a susceptible individual in order to prevent or treat an infectious disease 27. Possible sources of NAbs for SARS-CoV-2 are preparations generated in certain animal hosts or can be manufactured in the laboratory 7,27. However, a faster strategy to provide NAbs consists of transferring human convalescent plasma (CP) from previously infected individuals that have developed high titer NAbs that target SARS-CoV-2 13,55. CP has been used as a strategy of passive immunotherapy in prevention and management of infectious diseases since early 20th century, from there, it has been used to stem outbreaks of viral diseases such as, poliomyelitis, measles, mumps, and influenza 27,56. In the outbreaks of SARS-CoV, MERS-CoV, and H1N1 pandemic in 2009, CP therapy was successfully used as a treatment with satisfactory efficacy and safety 5. Hence, obtention of NAbs by transferring CP against SARS-CoV-2 to infected patients may be effective to neutralize the virus and to interrupt the vicious cycle of excessive activation of the immune response in severe patients 7,24. This strategy is a relatively rapid approach to obtain the standardized agents that could control COVID-19 and prevent further infection 41.
The mechanisms of the human defense include the production of antibodies which interact with SARS-CoV-2 and neutralize its actions 23. SARS-CoV-2 elicits a robust B cell response, as evidenced by the rapid and near-universal detection of virus specific IgM, IgG and IgA, and neutralizing IgG antibodies in the days following infection 13. A study reported that on day 5 after treatment, an increase of viral antibodies can be seen in nearly all patients, IgM positive rate increased to 81%, whereas IgG positive rate increased to 100%, which was considered as a transition from earlier to later period of infection 30.
Among all the major structural proteins, S protein is involved in receptor recognition as well as virus attachment and entry 57. The most important potential therapeutic for the treatment of SARS‐CoV-2 infection is to block the binding of S protein to ACE2 58. The S protein is an important target for NAbs to block binding and fusion of SARS-CoV-2 into the host cells 41. NAbs that target the S protein may provide temporary passive immunity to the disease 29. In the outbreak of SARS-CoV and MERS-CoV was discovered that NAbs bind to S1 subunit receptor-binding domain (S1-RBD) and S2 subunit of the S protein, thus inhibiting their entry, limiting viral amplification 56. Specific NAbs against RBD in S protein or that binds to ACE2 could effectively block the virus entry 7. One of the initial in-vitro studies that aimed to clone NAbs reported high titers of IgG antibodies against SARS-CoV-2 S protein, S1 subunit and its receptor-binding domain in most recovered patients 23.
Patients who have recovered from COVID-19 with a high NAbs titer may be a valuable donor source of CP 5. Currently, subjects between 18 and 65 without infectious symptomatology and a negative test for COVID-19 after 14 days of recovery are considered CP donors. Test must be repeated 48 hrs later of donation. To obtain CP, apheresis is the recommended procedure, which consists of a continuous centrifugation of blood from donor to allow a selective collection plasma 56. Clinical findings of transferring CP suggest that NAbs may improve the severity of the patients. In a clinical trial, Duan et al., reported that all symptoms, especially fever, cough, shortness of breath and chest pain, disappeared in all severe COVID-19 patients who received one dose of 200 mL CP within 1 to 3 days upon transfusion 5.
Nonetheless, the administration of CP carries risks, which are associated with the transfer of inadvertent infection or another infectious disease agent from the blood of the donor (27). The production of CP must be free of any infection. The process requires high quality standards in order to detect infectious agent such as, human immunodeficiency virus (HIV), hepatitis B, hepatitis C, syphilis and/or Trypanosoma cruzi (if living in an endemic area) (56). Another risk related to administration of CP is the reactions to serum constituents, including immunological reactions such as, serum sickness 27.
Some drugs previously tested in SARS-CoV are also a potencial treatment option to block the binding of SARS-CoV-2 to host cells. Emodin, an anthraquinone, could significantly block the interaction between the S protein and ACE2 by competing for the binding site of S protein in SARS-CoV. In addition, promazine, a phenothiazine class of antipsychotics, exhibited potent inhibition of the binding of S protein of SARS-CoV to ACE2. Therefore, emodin and promazine could be used as alternative choices for the treatment of COVID‐19 58.
Inhibiting the TMPRSS2
TMPRSS2 is responsible for priming and activating the S protein of SARS-CoV-2 allowing the entry of the virus into the host cells 57. One pathway for the development of therapeutics against SARS- CoV-2 is to block TMPRSS2 with a serine protease inhibitor 39. Nafamostat mesylate and camostat mesylate are known inhibitors of TMPRSS2. Currently, are approved in several countries to treat other conditions, while there are no clinical trials specifically testing these drugs against SARS-CoV-2 15. Although, camostat mesylate has been shown to partially block SARS-CoV-2 S protein-driven entry into Caco-2 and Vero-TMPRSS2 cells 59.
Blocking the Hemagglutinin-esterase (HE)
Arbidol, an indol derivate with broad-spectrum antiviral activity approved by Russia and China, is used for prophylaxis and treatment of influenza and respiratory viral infections 29,57. Arbidol mechanisms consist by blocking viral fusion to targeting HE, the major glycoprotein on the surface of influenza virus, preventing the fusion of the viral membrane with the endosome after endocytosis 28,57. Arbidol has demonstrated activity against several viruses including SARS. Currently, is being evaluated for the treatment of COVID-19 in several studies in China and is currently a recommended treatment option for COVID-19 there. In phase IV clinical trials, arbidol is provided to COVID-19 patients as an empirical therapy 28,29.
Inhibiting the RNA-dependent RNA polymerase (RdRp)
In CoVs, the RdRp catalyzes the synthesis of viral RNA, making it essential for viral replication and a prime target for antiviral inhibitors 13. Nucleoside analogs available for HIV and respiratory viruses may have a therapeutic role in blocking RNA synthesis by targeting the RdRp found in SARS-CoV-2 28.
Remdesivir
Among several potential drugs tested for efficacy in treatment of SARS-CoV-2 infection, remdesivir is considered as the most promising antiviral drug and shown to be the most hopeful anti-viral therapeutic for the treatment of COVID-19 30,60. Remdesivir is a 1’-cyano-substituted adenosine analog prodrug metabolized in cells and tissues to an active nucleoside triphosphate that inhibits viral RdRp by binding to RNA strands and preventing additional nucleotides from being added 13,28. Therefore, remdesivir it can incorporate into nascent viral RNA resulting in premature termination of the viral RNA chain and consequently halts the replication of the viral genoma 57.
Remdesivir was originally developed as a drug candidate for use in Ebola virus infection and has demonstrated broad-spectrum antiviral activity against filoviruses, paramyxoviruses, pneumoviruses, and pathogenic coronaviruses, including SARS-CoV and MERS-CoV 29,59. The first COVID-19 patient in the USA was successfully treated with remdesivir for the progression of pneumonia on day 7 of hospitalization without any noticeable adverse effect, and the patient’s condition improved on the 8th day in January, 2020 18,60,61. Remdesivir is now being tested in multiple trials in different countries, including two randomized phase III trials in China 18,29. Ongoing clinical trials are testing remdesivir with a dose of 200 mg intravenously followed by 100 mg/day intravenously in adult COVID-19 patients for 5 to 10 days 28,60.
Favipiravir
Similar to remdesivir, favipiravir, a prodrug licensed in Japan for the treatment of influenza, is another potential agent due to its activity against a wide spectrum of RNA viruses including CoVs 28. Favipiravir acts as a RdRp inhibitor by structurally resembling the endogenous guanine 57. In many RNA viruses, favipiravir is recognized as a competitive substrate of RdRp after to being converted into an active phosphoribosylated form, inhibiting the efficacy of viral replication 57.
Favipiravir is also under consideration for randomized clinical trials for COVID-19. In March 2020, the National Medical Products Administration of China approved favipiravir as the first anti-COVID-19 drug in China due to its efficacy and minimal side effects demonstrated in clinical trials 60.
Ribavirin
Ribavirin is a guanosine analogue antiviral prodrug that elicits its antiviral affect through inhibition of viral RNA synthesis 28,60. Ribavirin undergoes metabolic conversion in the liver, after which its metabolic structure functions as a purine guanosine analog which is incorporates into viral RNA sequences and prohibit the subsequent addition of nucleoside analogs, thereby preventing the synthesis of RNA 28.
Ribavirin has been used to treat several viral infections such as, hepatitis C, respiratory syncytial virus (RSV), and some viral hemorrhagic fevers. In most cases, ribavirin is combined with IFN 59. In SARS-CoV, the in vitro antiviral activity of ribavirin was estimated to be at a concentration of 50 mg/mL. However, it has the undesirable side effect of reducing hemoglobin, which is harmful to patients with respiratory distress (60). Molecular models indicate that ribavirin could effectively inhibit the RdRp of SARS-CoV-2. However, further studies are necessary in order to explore the efficacy of ribavirin for COVID-19 62.
Inhibiting the Protease Enzymes of SARS-CoV-2
Protease inhibitors are important agents in the contemporary treatment of patients with chronic HIV infection. In the Coronaviridae family, the targets of protease inhibitors are 3CLpro and papain-like proteases (PLPs) 60. 3CLpro and PLPs are CoVs encoded proteins that have an essential function for viral replication and additional function for inhibition of host innate immune responses. Targeting 3CLpro and PLPs are attractive options for the treatment of diseases caused by CoVs 58 The currently available HIV protease inhibitors have displayed some in vitro activity against the 3CLpro found in SARS-CoV and MERS-CoV 28.
Inhibiting the 3-chymotrypsin-like protease (3CLpro)
Lopinavir/ritonavir (LPV/r)
Lopinavir (LPV) is an aspartic acid protease inhibitor and a booster used for the treatment of HIV infection (30). LPV is co-formulated with ritonavir (LPV/r) to boost the pharmacokinetic activity and half-life of LPV through inhibition of Cytochrome P450 (CYP450). LPV/r provides potent and sustained viral load reductions in patients with HIV, which has concerted interest in assessing its efficacy in other viruses that function with similar machinery 28. Exist theoretical evidence that LPV/r also inhibit the 3CLpro protease encode by CoVs 57.
Based on findings in laboratory experiments, LPV/r could be used to treat the COVID-19 infections 43. Although, there are positive reports suggesting that LPV/r, have some clinical efficacy against SARS-CoV-2 29, a clinical trial conducted in patients with mild and moderate COVID-19, LPV/r showed little benefit for improving the clinical outcome. In another trial performed on patients with severe COVID-19, no benefits of LPV/r beyond standard care were observed 57. On July 4, 2020, WHO accepted the recommendation from the Solidarity Trial’s International Steering Committee to discontinue the clinical trials of LPV/r, since, these interim trial results show that LPV/r produce little or no reduction in the mortality of hospitalized COVID-19 patients when compared to standard of care 63.
An alternative drug corresponds to cinanserin, a serotonin receptor antagonist, which was a promising inhibitor of replication of SARS‐CoV due to 3CLpro inhibition. Therefore, cinanserin could be an option to evaluate the inhibition of 3CLpro in SARS-CoV-2 58.
Inhibiting the Viral papain-like protease (PLpro)
PLPs of human CoVs is a novel viral cysteine deubiquitinating enzymes (DUBs) which remove ubiquitin from target proteins and alter cellular pathways important for infection 64. In addition, PLPs are an IFN antagonist for inhibition of host innate antiviral immune response 58. Most CoVs can encode for up to two PLPs, however, MERS-CoV, SARS-CoV and SARS-CoV-2 encode for a single viral papain-like protease named PLpro 65. Studies have revealed that deubiquitinase activity of PLpro is tied to the pathogenesis of these CoVs. Therefore, the intrinsic activities of DUB and deISGylating of PLpro are believed to contribute to the pathogenesis of infection and to antagonize the activation of the host cell innate immune response by its downregulation of the IFN response 64,65.
In the ubiquinited system, the addition of ubiquitin to target proteins serves as a highly specific destruction tag, where ubiquitinated proteins are traced to the proteasome for degradation. The ISGylation consist in attach a small ubiquitin-like peptide (UBL) called Interferon-stimulated gene 15 (ISG15) covalently to proteins. ISG15 is a protein strongly induced by type I IFNs. Viral infection also strongly induces ISG15 because one of its major host responses is the production of type I IFNs. ISGylation can antagonize the formation of viral molecular complexes required for replication and activate molecules involved in immune pathways. Several proteins that are involved in antiviral signaling pathways, including RIG-I, MDA-5, Mx1, PKR, STAT1, and JAK1, have been identified as target proteins for ISGylation 64. Furthermore, the production of cytokines, chemokines, and other IFN-stimulated gene products with antiviral properties during infection can be regulated by ISGylation 65. A reverse pathway consists in deISGylating involving the removal of ISG15. deISGylating activity is mediated by the action of DUBs that catalyze the removal of ISG15 from tagged proteins. This action is necessary for viral replication and the evasion of host responses. PLPs specifically use their DUB activity to interfere with proteins that mediate signaling of viral infection, leading to a dysregulation of the immune pathways such as, NF-kB and IRF3 pathways, that in turn, results in a decrease in the antiviral response 64.
During viral infection, the recognition of the viruses triggers the activation of several signaling pathways and activation of transcription factors, such as, nuclear factor kappa B (NF-κB) and interferon regulatory factor 3 (IRF3). NF-κB stimulate the expression of several pro-inflammatory proteins, such as, tumor necrosis factor (TNF), cytokines (IL-1, IL-6 and IL-12) and chemokines (CCL2 and CXCL8) 52. IRF3 stimulate the production of type I Interferons (INF-α and INF-β) 2,35. In SARS-CoV, PLpro has shown that deubiquitinating and deISGylating activity inhibits the production of cytokines and chemokines and interferes with the host IRF3 and NF-kB antiviral pathways and modulates the host immune response. Inhibition of PLpro impacts virus replication via deficient viral protein processing and may also affect remote PLpro activities, including deubiquitination, de-ISGylation, and innate, anti-host immunity reactions. Inhibition of PLpro affects virus replication and its remote activities, including deubiquitination, de-ISGylation, and innate, anti-host immunity reactions 66. PLpro of SARS-CoV-2 also harbors deubiquitinating and deISGylating activities 65.
Naphthalene Based PLpro Inhibitors
Naphthalene based PLpro inhibitors have been shown to be effective at halting SARS-CoV-2 PLpro activity as well as SARS-CoV-2 replication. Naphthalene based PLpro inhibitors can serve as a jumping-off point for SARS-CoV-2 antiviral development. GRL-0617, a selective and competitive noncovalent inhibitor of SARS-CoV PLpro could be option to inhibit SARS-CoV-2 PLpro 65.
Disulfiram
A recent report demonstrated that disulfiram, a clinically approved alcohol abuse antagonist due to irreversible inhibition of aldehyde dehydrogenase (ALDH), is an inhibitor of the MERS-CoV and SARS-CoV PLPs, through a non-competitive and competitive mechanisms, respectively 64,67. Disulfiram has the capacity to covalently block the catalytic Cys residues of many enzymes, and conceivable, PLpro of SARS-CoV-2 67. Recent findings in vitro, together with a relatively good risk/benefit ratio, make disulfiram an evaluable drug against COVID-19. However, as a hepatic acetaldehyde dehydrogenase inhibitor, its pharmacological interactions must be considered 64.
Therapeutics Against Cytokine Storm Syndrome
Based on accumulated evidence, a subgroup of patients with severe COVID-19 might develop a cytokine storm syndrome 68. As opposed to directly targeting viral replication, other therapeutic approaches can be aimed at modulating the innate immune system or inhibit cytokines that are upregulated during viral replication to dampen the physiologic response to disease 28. In this context, the inhibition of this immunological pathway may help to control excessive cytokine production and may help in preventing further injury 56,69. In this way, treatment of cytokine storm syndrome with approved therapies with proven safety profiles has a strong theoretical rationale to address the immediate need to reduce mortality 68.
Blocking the Interleukin-6 (IL-6) Pathway
IL-6, TNF- α, and IL-1 are the most important pro-inflammatory cytokines in the human body. However, accumulated IL-6 is a predictive factor of poor prognosis in patients with ARDS 57. IL-6 is a widely used biomarker of inflammation. Elevated circulating IL-6 is associated with higher mortality in patients with community acquired pneumonia. IL-6 is also implicated in driving an excessive inflammatory response in the lungs of COVID-19 patients 34. Recently, it has also been reported that the elevated IL-6 is strongly associated with the need for mechanical ventilation 57. Consequently, a progressive rise in IL-6 may be a clinical indicator for the deterioration of COVID-19 30.
Several strategies are being evaluated to blunt IL-6 driven inflammation 34. The specific blockade of IL-6-regulated signaling pathways represents a promising approach in order to attenuate the damage associated with inflammation 57,70. Interleukin-6 receptor (IL-6R) has two forms: membrane-bound interleukin-6 receptor (mIL-6R) and soluble interleukin-6 receptor (sIL-6R). IL-6 binds to sIL-6R to form a complex, which then binds to glycoprotein 130 (gp130) on the cell membrane to complete trans signal transduction and play a proinflammatory role 71. Tocilizumab is a humanized monoclonal antibody which can specifically bind sIL-6R or mIL-6R and inhibit signal transduction 30. Tocilizumab has been widely used in the treatment of chimeric antigen receptor (CAR)-T cell-induced cytokine release syndrome (CRS), rheumatoid arthritis, adult-onset Still’s disease, giant cell arteritis and polyarticular or systemic juvenile idiopathic arthritis 28,30,72. A multicentre, randomized controlled trial of tocilizumab has been approved in COVID-19 patients with associated pneumonia and elevated IL-6 in China. Preliminary data reported that fever subsided in all patients within the first day of receiving tocilizumab and oxygen requirements were reduced in 75% of the patients 68,72.
Corticosteroids
Corticosteroids are an adjuvant therapy for ARDS and cytokine storm syndrome due to their anti-inflammatory activity. However, the broad immunosuppression mediated by corticosteroids does raise the possibility that treatment could interfere with the development of a proper immune response against the virus 13. According to current WHO interim guidance on COVID-19 management, corticosteroids were not recommended as routine therapy unless alternative indications are present, because possible harms and higher risk of mortality attributed to corticosteroids therapy have been reported by studies on other CoVs and influenza 28. Corticosteroids might be used as an adjunct for treating severe forms of the disease and should only be considered if required for an alternative indication such as, septic shock or bronchoconstriction 28,59. The use of short-term corticosteroid therapy could be considered in patients who present deterioration in oxygenation index, or rapid progression of radiological findings, or excessive activation of immune responses within 10 days of illness onset 30. The administration of methylprednisolone would help suppress the unwanted immune reactions, but their use is controversial. Nevertheless, studies have been initiated to explore its effectiveness and safety 57. A clinical trial is actively recruiting to evaluate the efficacy and safety of methylprednisolone with a dose of 1 mg/kg/day for a duration of 7 days in noncritically ill patients to further identify the risks and benefits of therapy 38. A Retrospective cohort study of 201 patients with confirmed COVID-19 in China, 44 patients developed ARDS. Among patients with ARDS, methylprednisolone decreased the risk of death (HR, 0.38; 95% CI, 0.20-0.72) 73. In this sense, treatment with methylprednisolone may be beneficial in severe and critical cases. For severe cases, Zhou et al., suggest start at a dose of 40 to 80 mg/day methylprednisolone and slowly taper over 7 to 10 days, and some suggested for a shorter period of 3 to 5 days. For critically ill cases, a starting dose of 80 to 160 mg/day methylprednisolone, following a slow withdrawal within 7 to 10 days is considered 30.
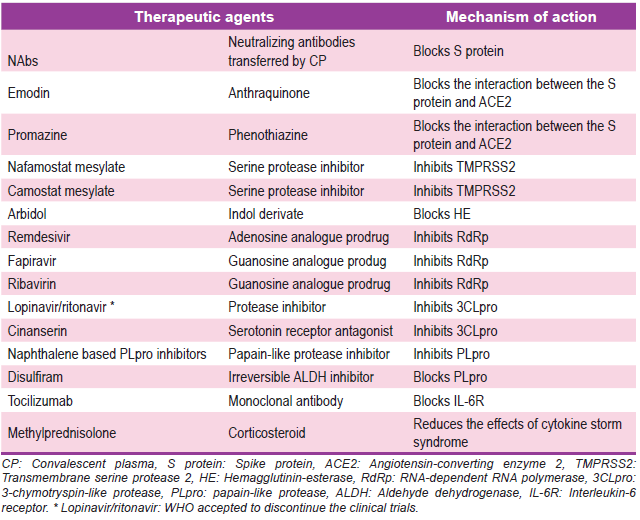
Finally, a summary of the potential therapeutic agents against SARS-CoV-2 is shown in (Table 1).
CONCLUSION
The outbreak of the novel coronavirus pandemic leading to COVID-19 disease has dramatically accelerated research into effective drugs and a vaccine to treat this disease. Currently, the main clinical treatment consists of symptomatic management and oxygen therapy, with mechanical ventilation for patients with respiratory failure. Thus far, there is no approved treatment or vaccine for SARS-CoV-2. In the absence of a specific treatment, the strategy options are based on previous experience in treating other viral infections. The potential pharmacologic therapies against SARS-CoV-2 target its viral structure and the mechanisms of viral life cycle.
SARS-CoV-2 expresses S protein on its outer surface that facilitate attachment on to host cells via ACE2. Among immunotherapy approaches to block the virus attachment or entry, NAbs that target the S protein, could provide an effective treatment due to their specificity. A high titer of NAbs can faster isolated and transferred from the convalescent plasma (CP) of the previously infected patients, providing temporary passive immunity to the disease.
For the replication of its genome, SARS-CoV-2 recruits viral protease enzymes such as, 3CLpro and PLpro, and non-structural proteins such as, RdRp. The use of antivirals may inhibit these enzymes and prevent a successful virus cycle of infection. Among potential drugs tested for the treatment of SARS-CoV-2, remdesivir is considered as the most promising antiviral drug. However, the efficacy of remdesivir must be determinated from the numerous clinical trials that are still ongoing.
Other therapeutic approaches can be aimed at treating the severity of cytokine storm syndrome. In elderly or with comorbidities patients, a rapid viral replication may cause tissue injury, leading to induce the exaggerated production of pro-inflammatory cytokines such as IL-6. An elevated IL-6 is associated as a clinical warning indicator for the deterioration of COVID-19. Tocilizumab, a monoclonal antibody used in the treatment of autoimmune diseases, can specifically bind IL-6 receptors, and inhibit signal transduction. Blocking IL-6 receptors could mitigate cytokine storm syndrome related symptoms in severe COVID-19 patients. In addition, corticosteroids are an adjuvant therapy for cytokine storm syndrome. However, the current WHO interim guidance on the management of COVID-19 strongly recommends against the use of corticosteroids unless indicated for another reason. Patients who present excessive activation of immune responses within 10 days of illness onset could be considered for the use corticosteroids. In this sense, treatment with methylprednisolone may be an option in severe and critical cases.
Conflict of Interest Statement
The author declares no conflict of interest.